But when I am tried to open it, '.h5' was not in the list of supported. I wrote a data file using HDF for fortran and generated a file (filename.h5). To avoid the problem of manually customizing the GUI for each application that is to be steered, we make use of XML templates that describe outputs from the simulation (and inputs back to it) to automatically generate GUI controls for manipulation of the simulation. In all of Paraview manuals it says that it is compatible with HDF5 file format. This allows not only simple parameter changes, but complete remeshing of grids, or operations involving regeneration of field values over the entire domain. By making use of a distributed shared memory file, one may read data from the simulation, modify it using ParaView pipelines, write it back, to be reused by the simulation (or vice versa). Each MPI job may use different core counts or hardware configurations, allowing fine tuning of the amount of resources dedicated to each part of the workload. Our implementation using ParaView as the interface, allows a flexible combination of parallel simulation, concurrent parallel analysis, and GUI client, either on the same or separate machines. We achieve this by replacing the IO layer in the HDF5 library with a custom driver which transfers data in parallel between simulation and analysis. 
Writer.Interfacing a GUI driven visualization/analysis package to an HPC application enables a supercomputer to be used as an interactive instrument. Writer.write_points_cells(mesh.points, mesh.cells) Mesh = meshio.read(f'mesh/mesh.xdmf') as writer: Sample data is available at MPCDF DataShare (password: viz)Ĭurrently, I convert the VTK mesh files in mesh and the data in the snaps files into XDMF+HDF5 files via the following simple script. 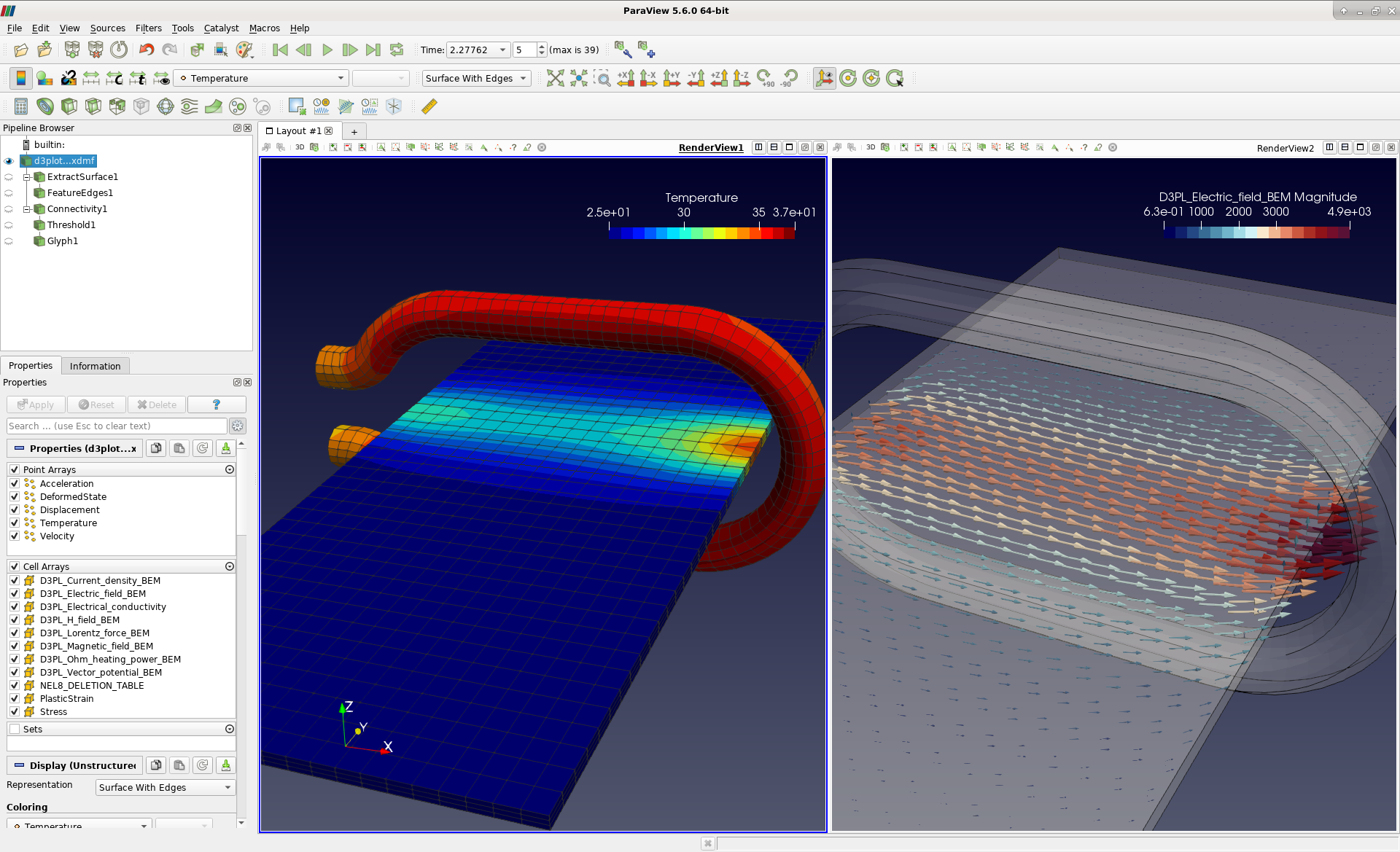
Is there a better way to handle the time dimension (and to work around the hyperslab issue) in XDMF (preferably using Xdmf3ReaderS to spatially parallelise)? Or is there another “light+heavy” format which is easier to get a time series into?Īny help/pointers would be greatly appreciated! Sample data However, can these formats be used as a ‘light’ data descriptor – I don’t see a way to point to existing data files with these? I think I’d need to reformat my data file, or at least copy a EXODUS II/CGNS header into my HDF5 files (the latter option is possible, but I don’t see documentation on how to do this for time series)Īs such, I’m not sure how I should go about reading my data files. As such, formats like EXODUS II or CGNS are recommended over XDMF. PDF We present a framework for interfacing an arbitrary HPC simulation code with an interactive ParaView session using the HDF5 parallel IO library as. Finally, from other posts, it sounds like the XDMF project is no longer developed. Package: paraview Maintainer for paraview is Debian Science Team <> Source for paraview is src:paraview (PTS, buildd, popcon).Additionally, the XDMF reader (all 3 versions) only loads the grid if I add a hyperslab: seems possibly related to, see below paraview: cant load hdf5 file: vtkSISourceProxy: Failed to create vtkXdmfReader.Now I am trying to reverse engineer the file from my tool. I initially created a VTK-file from my model and used Paraview -> File -> Save Data -> Xdmf Data File (.xmf) to create a working combination of light and heavy data in Xdmf/HDF5. For example: with h5py.File(root.withsuffix(.h5)) as file, xh.Grid(root.withsuffix(.xdmf)) as xdmf: xdmf +. Apparently improved handling of time-data was under development in XDMF, but it doesn’t appear to be there yet (if ever) Hi, I am trying to write a Xdmf-file from my tool for visualization in ParaView or VisIt using the HDF5 format. Module to write XDMF files for HDF5 files. Additionally, each time a new time-point is added, I’d need to modify the XML. I have tried to use hyperslab-indexing and a temporal collection to point to each time point in each file, but then I’d need to make N times T entries in the XDMF file (i.e. The natural solution would be something like XDMF, where some ‘light’ description file is used to describe the interface to the ‘heavy’ HDF-5 files. However, since the files are large, I would like to interface with them directly rather than copying them into a file format like VTK, and since we have a pre-existing library of analysis tools I can’t change the format. Currently, I can plot the data by using meshio to copy the fields into a format understood by Paraview – this was used to make the above figure.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
